yunha[at]mit[dot]edu

About me
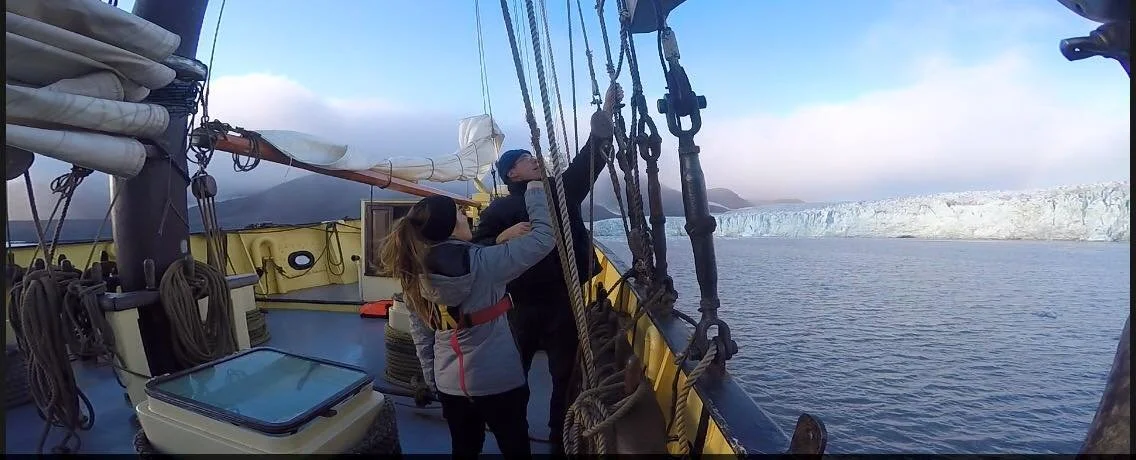
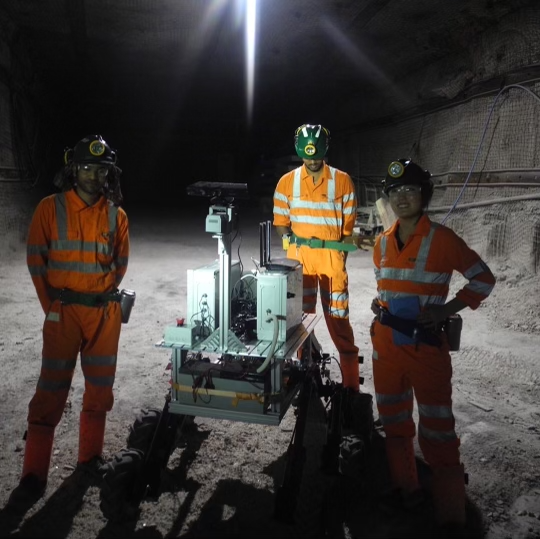
I am an Assistant Professor at MIT with a shared appointment between Biology, EECS and the Schwarzman College of Computing. I also serve as the co-founder and Chief Scientist at Tatta Bio, a scientific nonprofit dedicated to advancing genomic AI for biological discovery. I completed my Ph.D. in Biology from Harvard University and B.S. in Computer Science from Stanford University. My research interests span machine learning for sustainable biomanufacturing, microbial evolution, and open science.
I am looking for PhD students and postdocs to join my lab in Fall 2025, please email me if you are interested.
Research Interests
Microbial genomes encode the largest molecular, biochemical and functional diversity on Earth. My group’s research focuses on developing machine learning models and experimental approaches to discover and design novel biological functions. We integrate computation with expertise in evolution, ecology and biochemistry to characterize and harness the functional potential of microbes.
Machine learning approaches to discover and design microbial biochemistry. Microbes are the world’s best chemists. We develop machine learning approaches and datasets to systematically explore microbial chemical diversity.
Modeling and interpreting mechanisms of microbial evolution, ecology and function. AI is transforming how we conduct scientific inquiry. We build and interpret biological sequence models to discover mechanisms that underpin biological processes across scales.
Microbial applications for human and environmental health. From natural products, biomanufacturing to microbial therapies, we focus on research that addresses critical challenges.
Publications
For the most up to date list, please see my Google Scholar.
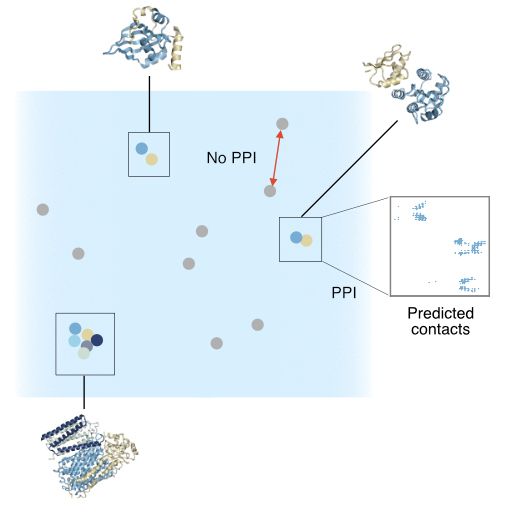
Andre Cornman, Matt Tranzillo, Nicolo G. Zulaybar, Imane Bouzit, Yunha Hwang, “Linear-time prediction of proteome-scale microbial protein interactions”, bioRxiv 2026.03.01.708874; doi: https://doi.org/10.64898/2026.03.01.708874
Nishant Jha, Joshua Kravitz, Jacob West-Roberts, Antonio Camargo, Simon Roux, Yunha Hwang, “Gaia: A Context-Aware Sequence Search and Discovery Tool for Microbial Proteins”, Science Advances, https://doi.org/10.1126/sciadv.adv5109 (2024)
Try out Gaia at https://seqhub.org

Andre Cornman, Jacob West-Roberts, Antonio P Camargo, Simon Roux, Martin Beracochea, Milot Mirdita, Sergey Ovchinnikov, Yunha Hwang , “The OMG dataset: An Open MetaGenomic corpus for mixed-modality genomic language modeling”, ICLR (2025)

Jacob West-Roberts, Joshua Kravitz, Nishant Jha, Andre Cornman, Yunha Hwang “Diverse Genomic Embedding Benchmark for functional evaluation across the tree of life”, BioRxiv, (2024)

Yunha Hwang, Andre Cornman, Elizabeth Kellogg, Sergey Ovchinnikov, Peter Girguis “Genomic language model predicts protein co-regulation and function” Nature Communications, (2024)

Yunha Hwang, Simon Roux, Clement Coclet, Sebastian Krause, Peter Girguis “Viruses interact with hosts that span distantly related microbial domains in dense hydrothermal mats.” Nature Microbiology, (2023)

Yunha Hwang, Peter Girguis “Differentiated evolutionary strategies of atlantic and pacific thaumarchaeal populations” mSystems, (2022)

Yunha Hwang, Dirk schulze-Makuch, Felix Arens, Johan Saenz, Panagiotis Adam, Christof Sager, Till Bornemann, Weishu Zhao, Ying Zhang, Alessandro Airo, Michael Schloter, Alexander Probst “Leave no stone unturned: Individually adapted xerotolerant Thaumarchaeota sheltered below the boulders of the Atacama Desert hyperarid core” Microbiome, (2021)

Yunha Hwang, Janina Rahlff, Dirk Schulze-Makuch, Michael Schloter, Alexander Probst “Diverse viruses carrying genes for microbial extremotolerance in the Atacama Desert hyperarid soil” mSystems, (2021)








